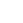The core principle of nanopore sequencing is to read sequence information by detecting the current changes caused by a single DNA or RNA molecule passing through the nanopore. However, the original electrical signal is continuous and complex, requiring efficient decoding algorithms to convert it into accurate base sequences. The decoding algorithm analyzes and processes the current signal to identify the characteristic signal pattern corresponding to each base, thereby achieving the conversion from electrical signals to base sequences. The quality of decoding algorithms directly affects the accuracy of sequencing results. High quality decoding algorithms can more accurately parse complex current signals, reduce misreading rates, and improve the accuracy of base recognition.

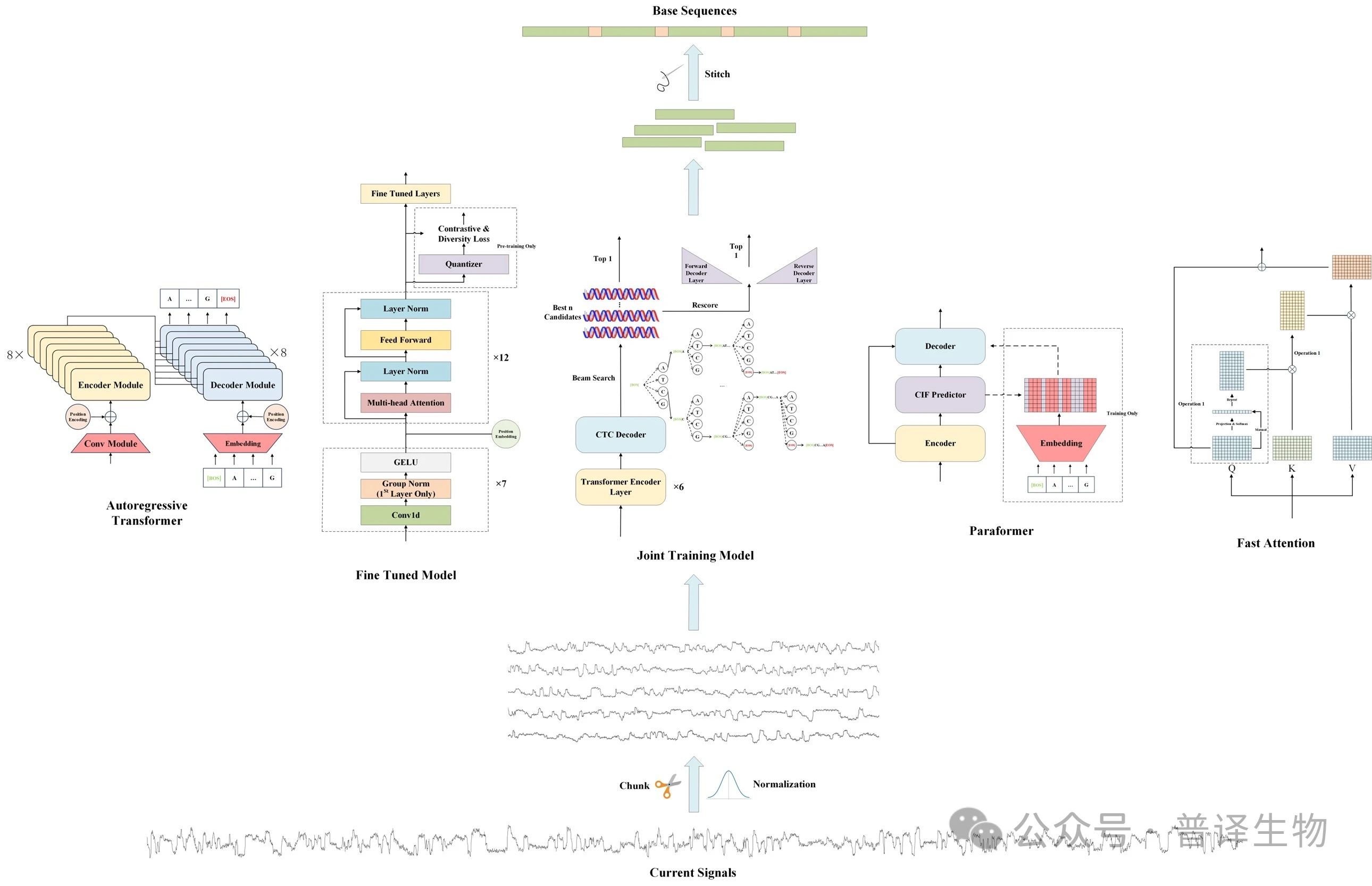
Figure 1. Working principle of BaseNet
On September 25, 2024, Polyseq Biotechnology and the Institute of Biophysics of the Chinese Academy of Sciences jointly published a research paper entitled "BaseNet: A Transformer Based Toolkit for Nanopore Sequencing Signal Decoding" online in the Computational and Structural Biotechnology Journal. This study proposes a nanopore sequencing signal decoding toolkit BaseNet (Figure 1) based on multiple latest Transformer algorithms, including:
1. Autoregressive Transformer based on Beam Search;
2. A non autoregressive Transformer trained using the Rescore mechanism and combined loss of CTC and AED;
3. Paraformer based on CIF predictor and GLM generator;
4. A large-scale pre trained model based on contrastive learning and diversity learning;
5. Fast attention with linear computational complexity.
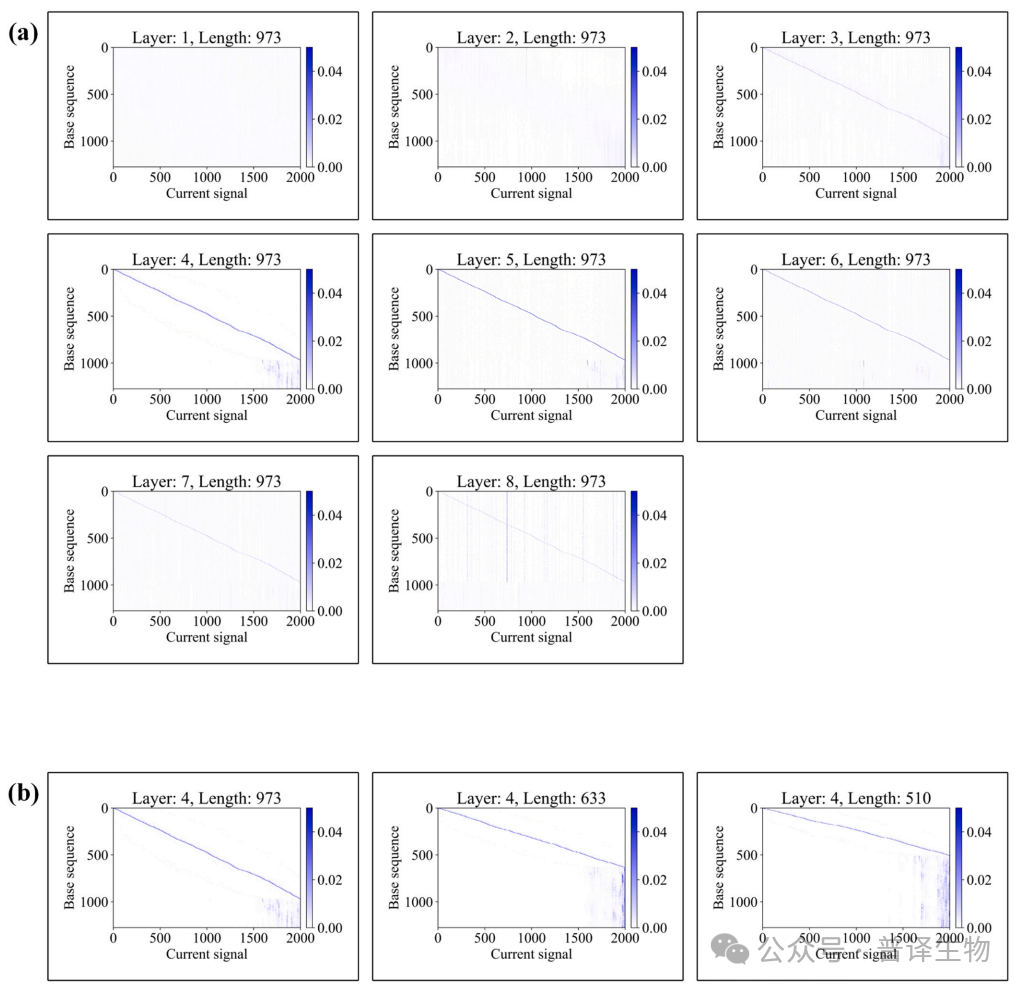
Figure 2. Visualization of Cross Attention Score
And exploration was conducted on the interpretability of the model, such as the fact that cross attention scores can map the alignment relationship between signals and sequences (Figure 2), and that speech waveforms and current signals share common "universal features" in the internal representation of the model (Figure 3).
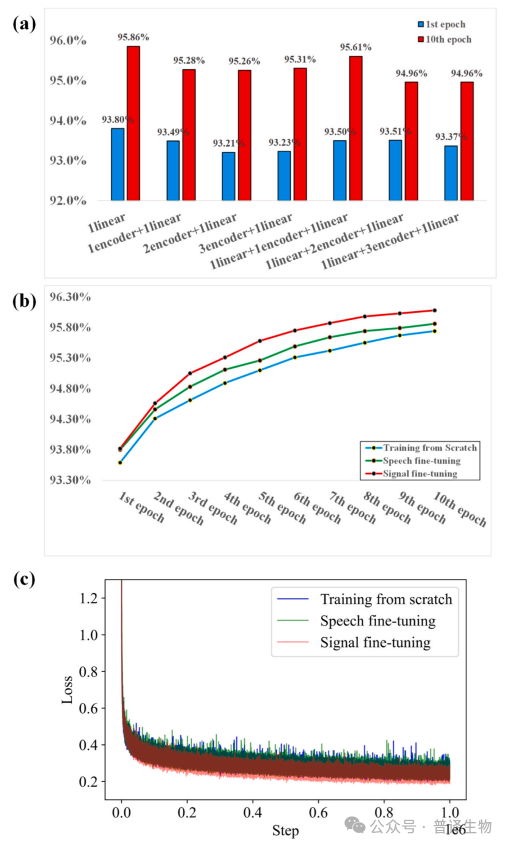
Figure 3. Performance Comparison of Different Large Models
The research team conducted rigorous performance evaluations on the same benchmark dataset, and found that the Fine tuned model included in BaseNet outperformed the latest Bonito CRF model, while the Joint CTC model and Fast CRF model performed better than SACall (Figure 4a). In addition, under long read conditions, the model developed by BaseNet exhibits superior performance (Figure 4b). The above test results indicate that BaseNet exhibits better decoding performance compared to other existing decoding algorithms.
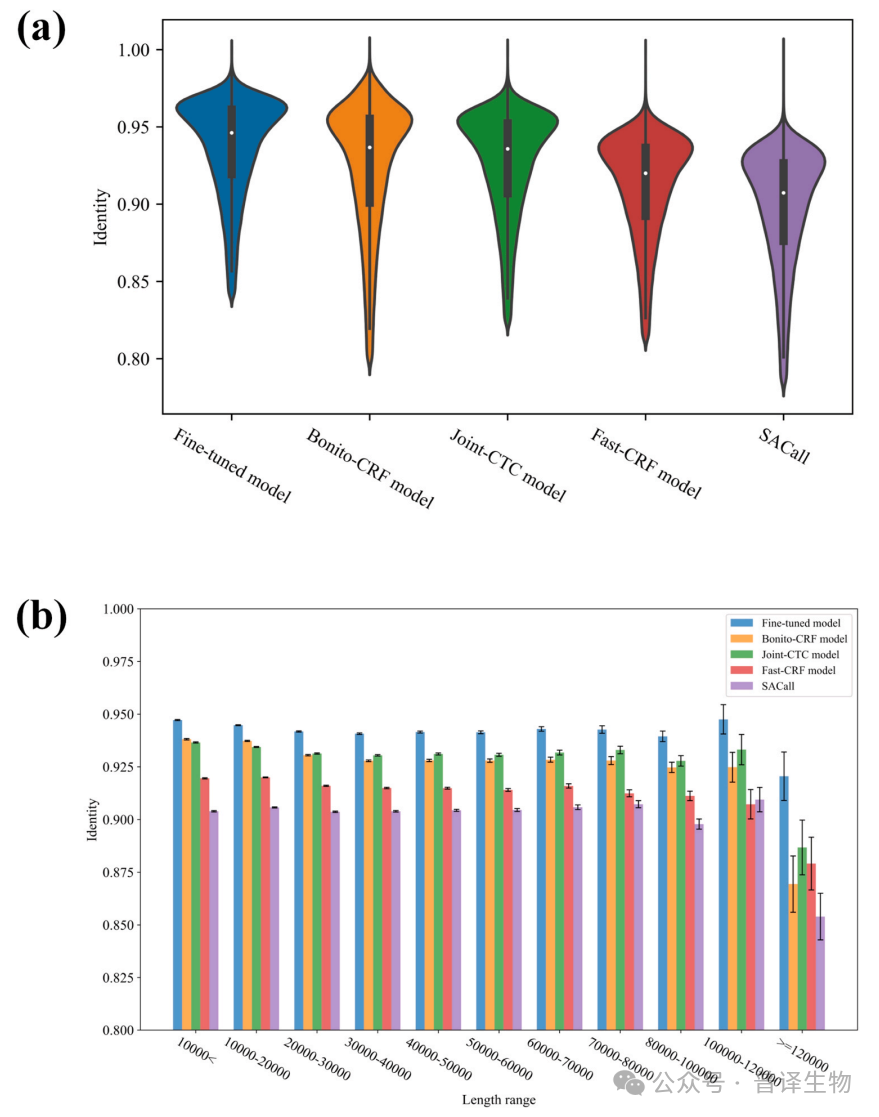
Figure 4. Performance comparison between BaseNet and existing Basecallers
Dr. Wang Daqian, General Manager of Polyseq Biotechnology, and Dr. Lou Jizhong, Chief Scientist, are the co corresponding authors of the paper. Li Qingwen, a joint doctoral student from Polyseq Biotechnology and Biophysics Institute, is the first author of the paper. Sun Chen from Polyseq Biotechnology participated in the research work.